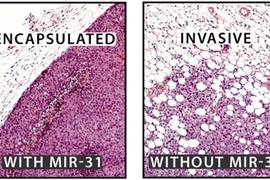
High levels of a tiny fragment of RNA appear to suppress the spread of breast cancer in mice, according to researchers at Whitehead Institute for Biomedical Research.
Measuring levels of this so-called microRNA, which is also associated with metastatic breast cancer in humans, may more accurately predict the likelihood of metastasis (which accounts for 90 percent of cancer-related deaths) and ultimately help determine patient outcomes.
In the study, reported in the June 12 issue of Cell, Scott Valastyan, a graduate student in Whitehead Member Robert Weinberg's laboratory, screened patient breast cancer samples for microRNAs with potential roles in metastasis. MicroRNAs are single strands of RNA about 21 to 23 nucleotides long. Within a cell, a single microRNA can fine-tune the expression of dozens of genes simultaneously. This capability could be particularly important in metastasis, a multistep process that could be influenced by a single microRNA at several points.
The screened samples were classified as either metastatic cancer or non-metastatic cancer. After analysis, the microRNA miR-31 stood out because of its inverse correlation with metastasis. In samples where a patient's original tumor had not metastasized, the cancer cells retained high levels of the microRNA. But where the tumor had metastasized, the cancer cells expressed lower levels of miR-31.
The functional role of miR-31 in metastasis regulation was then confirmed in mice. When Valastyan removed miR-31 from normally non-aggressive breast cancer cells and implanted those cells into mice, the cells formed highly aggressive tumors. Mice injected with the cancer cells lacking miR-31 had six to 10 times more cancer cells that metastasized to their lungs than did their counterparts implanted with unmodified cancer cells.
To see how increasing miR-31 levels could affect metastasis, Valastyan introduced miR-31 into breast cancer cells that readily metastasize. After injecting these altered cells into mice, the mice had as few as one-fortieth the metastases as mice injected with the unaltered cells.
Valastyan says that quantifying miR-31 levels in a patient's cancer cells could one day support a more accurate prognosis. Currently, breast cancers are divided into three major categories, two of which are typically associated with poor prognoses.
"This microRNA seems to be quite unique, in that it seems to provide some prognostic utility across these existing subclassifications [of cancers]," says Valastyan. A better-defined prognosis could help patients determine whether they might benefit from poorly tolerated cancer therapies.
In addition, miR-31 could be a useful target for cancer therapy. Weinberg, who is also a professor of biology at MIT, is cautiously optimistic. "At present, it's quite difficult to inhibit the action or promote the actions of a microRNA in a whole organism," he says, "but in the future, microRNAs like this one might prove to be very important in altering the clinical progression of a tumor or causing it to revert to a more benign state."
This research was supported by the National Institutes of Health (NIH), MIT Ludwig Center for Molecular Oncology, U.S. Department of Defense (DoD), Breast Cancer Research Foundation, Harvard Breast Cancer Specialized Program of Research Excellence (SPORE), and a DoD Breast Cancer Research Program (BCRP) Idea Award.